Regression with rbbnp
Xinyu Dai and Susanne M. Schennach
2026-02-14
regression.RmdThis article provides a comprehensive guide to conditional
expectation (regression) estimation using
biasBound_condExpectation().
The biasBound_condExpectation() Function
biasBound_condExpectation(
Y, # Response variable
X, # Predictor variable
x = NULL, # Evaluation points
h = NULL, # Bandwidth
h_method = "cv", # "cv" or "silverman"
alpha = 0.05, # Confidence level
kernel.fun = "Schennach2004"
)Basic Example
# Generate data: Y = f(X) + noise
set.seed(42)
X <- gen_sample_data(size = 800, dgp = "2_fold_uniform")
Y <- sin(2 * pi * X) + rnorm(800, sd = 0.3)
# Estimate conditional expectation E[Y|X]
fit <- biasBound_condExpectation(Y, X, h = 0.1, kernel.fun = "Schennach2004")
# View results
fit
#> Bias-Bounded Conditional Expectation Estimation
#>
#> Call:
#> biasBound_condExpectation(Y = Y, X = X, h = 0.1, kernel.fun = "Schennach2004")
#>
#> Sample size: n = 800
#> Bandwidth: h = 0.1000 (user-specified)
#> Kernel: Schennach2004
#>
#> Bias bound parameters:
#> A = 2.2646, r = 1.4677, B = 0.6769
#> bias bounds: b1x = 0.5229, byx = 0.5229
#>
#> Evaluation points: 100 (range: [0.0688, 1.9084])
#> Fitted values: E[Y|X] range [-1.1778, 1.2803]
#> Confidence level: 95%
#>
#> Use summary() for detailed statistics
#> Use plot() to visualize results
#> Use fitted() to extract fitted valuesUnderstanding the Output
# Key parameters
coef(fit)
#> A r B h
#> 2.2646049 1.4677318 0.6769206 0.1000000
# Detailed summary
summary(fit)
#> Summary: Bias-Bounded Conditional Expectation Estimation
#> ============================================================
#>
#> Call:
#> biasBound_condExpectation(Y = Y, X = X, h = 0.1, kernel.fun = "Schennach2004")
#>
#> Sample Information:
#> Sample size (n): 800
#> Bandwidth (h): 0.1000
#> Kernel function: Schennach2004
#>
#> Bias Bound Parameters:
#> A (amplitude): 2.2646
#> r (decay rate): 1.4677
#> B (Y bound): 0.6769
#> b1x (bias f(x)): 0.5229
#> byx (bias f_YX): 0.5229
#> Xi interval: [2.4522, 13.0417]
#>
#> Range Estimation:
#> Fitted values (E[Y|X]):
#> min Q1.25% median mean Q3.75% max
#> -1.1778 -0.7739 0.0083 0.0136 0.7794 1.2803
#>
#> Marginal density f(x):
#> min mean max
#> 0.0911 0.5333 0.9695
#>
#> Standard errors:
#> min mean max
#> 0.0078 0.0139 0.0193Visualizing Results
plot(fit)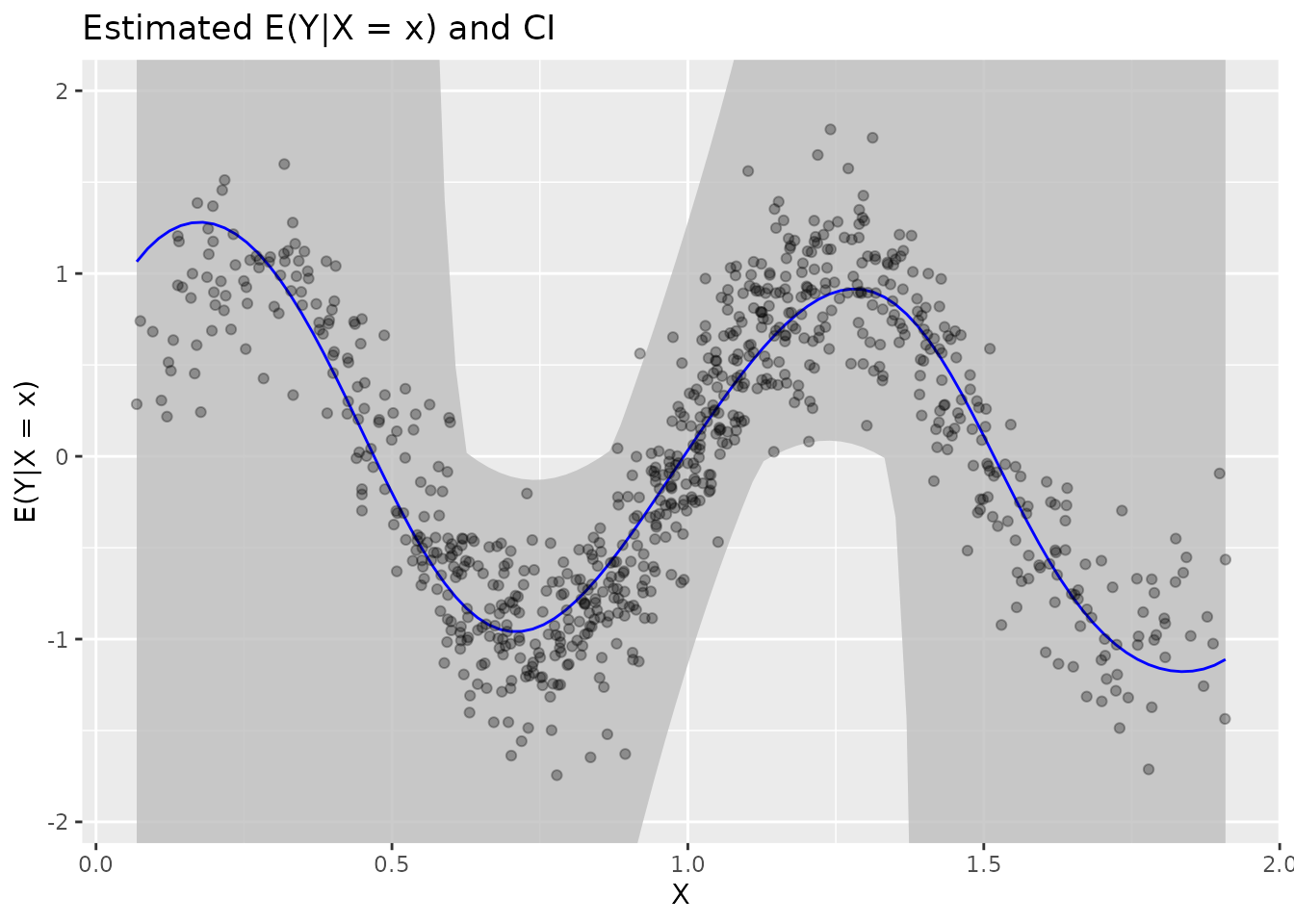
Plot elements:
| Element | Description |
|---|---|
| Blue line | Estimated |
| Grey band | 95% confidence interval |
| Black points | Original data |
Adding True Function
# True function for comparison
true_fn <- function(x) sin(2 * pi * x)
plot(fit) +
stat_function(fun = true_fn, aes(color = "True E[Y|X]"),
linetype = "dashed", linewidth = 1) +
scale_color_manual(values = c("True E[Y|X]" = "red")) +
labs(color = NULL) +
theme(legend.position = "top")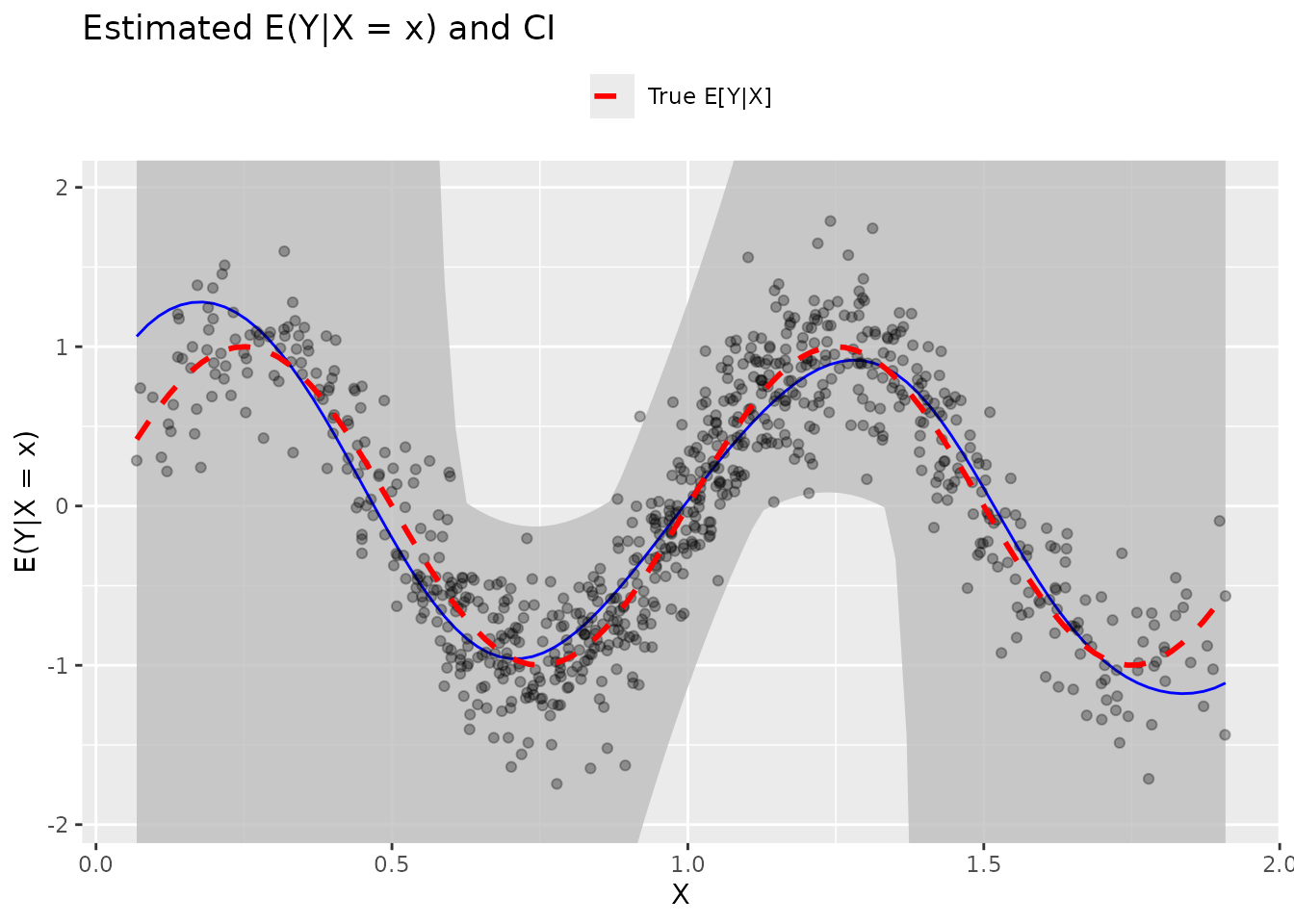
Extracting Fitted Values
# Get fitted values at evaluation points
predictions <- fitted(fit)
head(predictions, 10)
#> [1] 1.064898 1.136427 1.192210 1.233783 1.261980 1.277342 1.280253 1.270928
#> [9] 1.249450 1.215965
# Evaluation points
x_points <- fit$x
head(x_points)
#> [1] 0.06877618 0.08735791 0.10593964 0.12452137 0.14310310 0.16168483Confidence Intervals
In regions where the estimated marginal density
is very close to zero, the confidence interval may become
unbounded (i.e., contain -Inf or
Inf). This behavior is expected because the conditional
mean estimator is formed as a ratio involving
.
# Extract confidence intervals
ci <- confint(fit)
head(ci, 10)
#> lower upper
#> [1,] -Inf Inf
#> [2,] -Inf Inf
#> [3,] -Inf Inf
#> [4,] -Inf Inf
#> [5,] -Inf Inf
#> [6,] -Inf Inf
#> [7,] -Inf Inf
#> [8,] -Inf Inf
#> [9,] -Inf Inf
#> [10,] -Inf Inf
# Manual extraction for custom analysis
data.frame(
x = fit$x[1:6],
estimate = fitted(fit)[1:6],
lower = ci[1:6, "lower"],
upper = ci[1:6, "upper"]
)
#> x estimate lower upper
#> 1 0.06877618 1.064898 -Inf Inf
#> 2 0.08735791 1.136427 -Inf Inf
#> 3 0.10593964 1.192210 -Inf Inf
#> 4 0.12452137 1.233783 -Inf Inf
#> 5 0.14310310 1.261980 -Inf Inf
#> 6 0.16168483 1.277342 -Inf InfBandwidth Selection
Cross-Validation
fit_cv <- biasBound_condExpectation(Y, X, h = NULL, h_method = "cv")
cat("CV bandwidth:", coef(fit_cv)["h"])
#> CV bandwidth: 0.2313533Silverman’s Rule
fit_silv <- biasBound_condExpectation(Y, X, h = NULL, h_method = "silverman")
cat("Silverman bandwidth:", coef(fit_silv)["h"])
#> Silverman bandwidth: 0.1156766Real Data Example
Let’s use the built-in sample data:
# Load sample data
data(sample_data)
head(sample_data)
#> X Y
#> 1 1.0859914 2.828175
#> 2 1.6292714 2.014698
#> 3 1.2303920 3.350186
#> 4 1.0686912 1.953532
#> 5 0.8441481 1.391845
#> 6 0.8789330 1.954778
# Estimate regression
fit_real <- biasBound_condExpectation(
Y = sample_data$Y,
X = sample_data$X,
h = 0.1,
kernel.fun = "Schennach2004"
)
# Visualize
plot(fit_real) + ggtitle("Regression on Sample Data")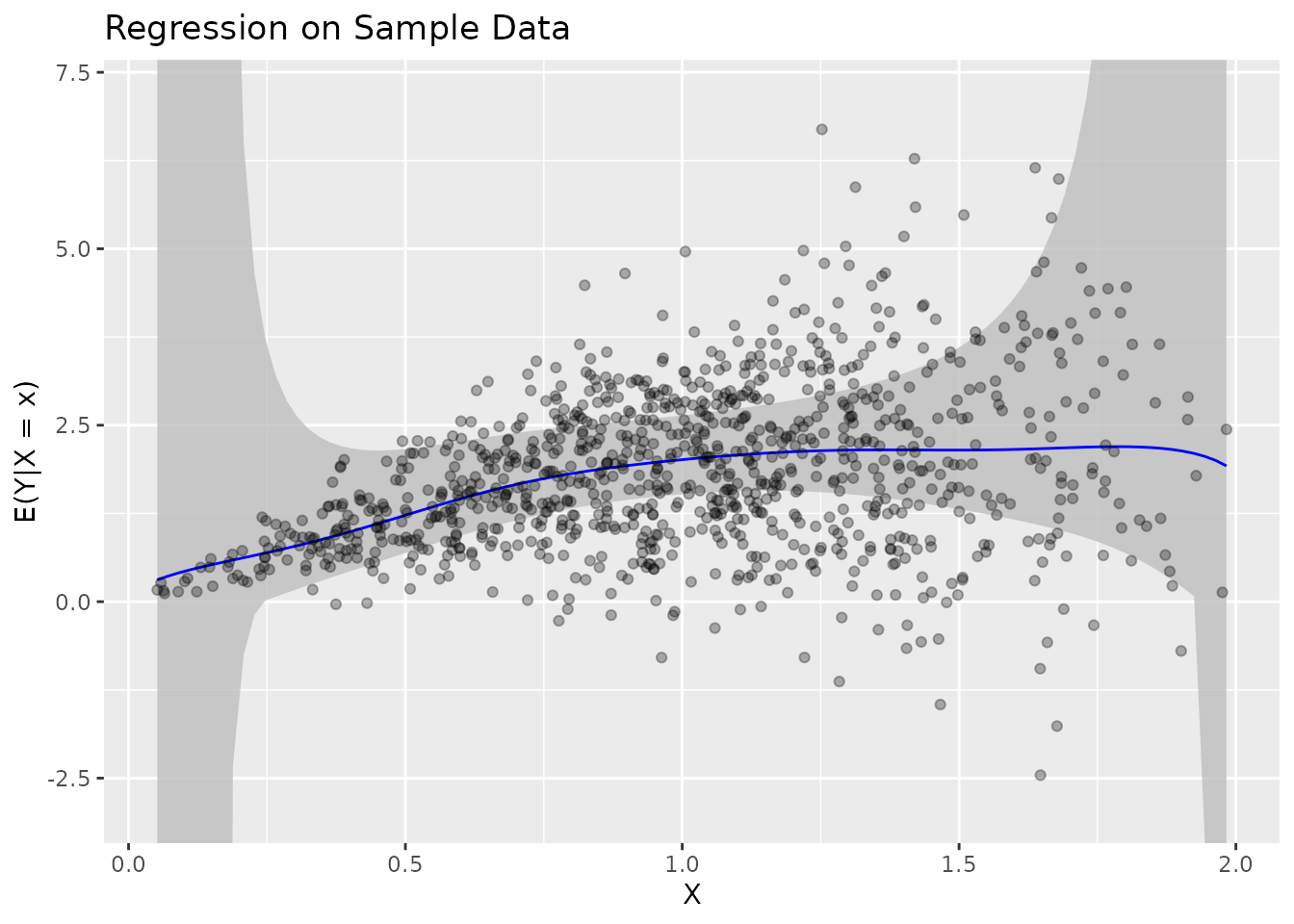
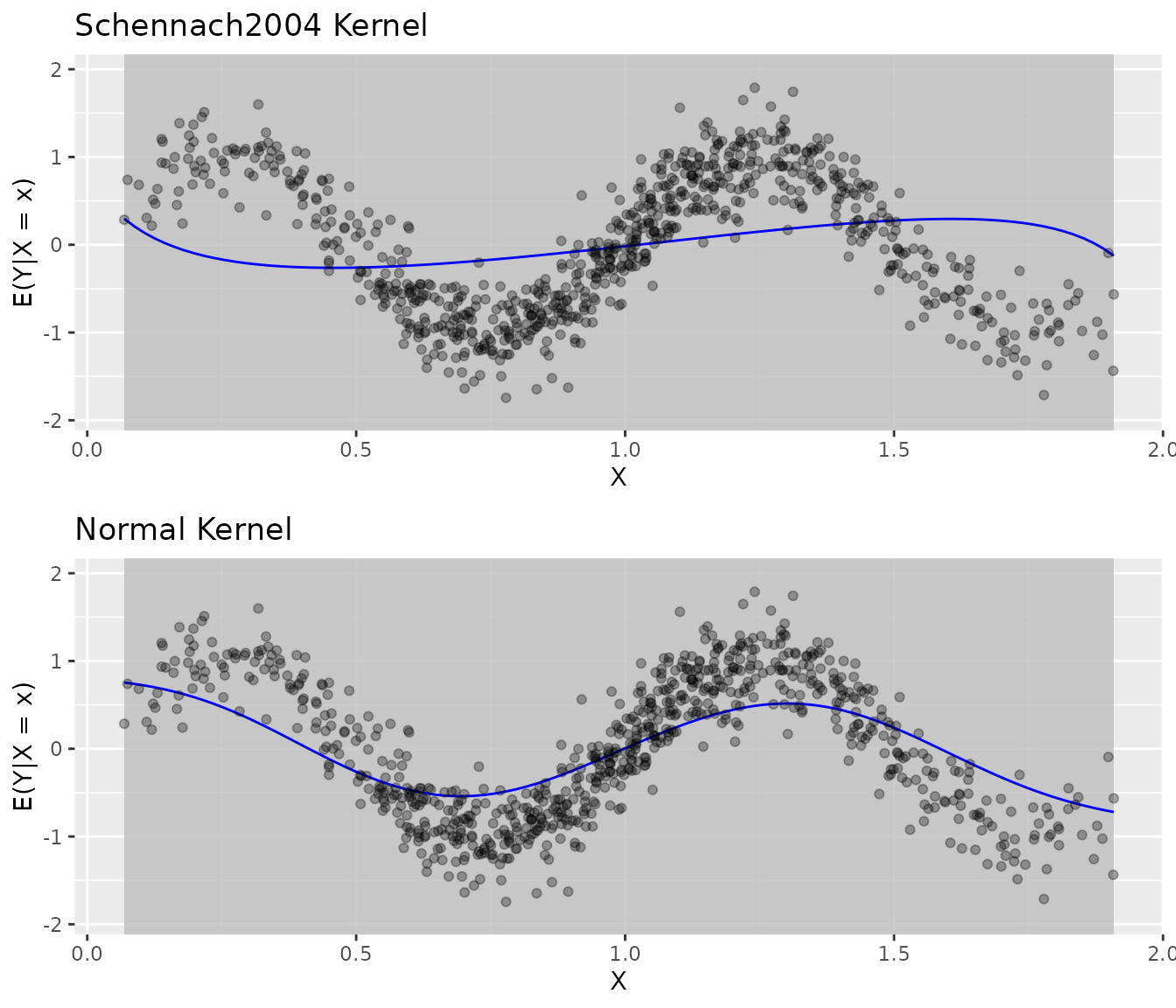
Comparing Kernel Functions
fit_sch <- biasBound_condExpectation(Y, X, kernel.fun = "Schennach2004")
fit_norm <- biasBound_condExpectation(Y, X, kernel.fun = "normal")
grid.arrange(
plot(fit_sch) + ggtitle("Schennach2004 Kernel"),
plot(fit_norm) + ggtitle("Normal Kernel"),
ncol = 1
)
Heteroscedastic Data
The method handles heteroscedastic errors:
# Generate heteroscedastic data
set.seed(123)
X_het <- gen_sample_data(size = 600, dgp = "2_fold_uniform")
Y_het <- X_het^2 + rnorm(600) * X_het # Variance increases with X
fit_het <- biasBound_condExpectation(Y_het, X_het, h = 0.1)
plot(fit_het) + ggtitle("Heteroscedastic Data")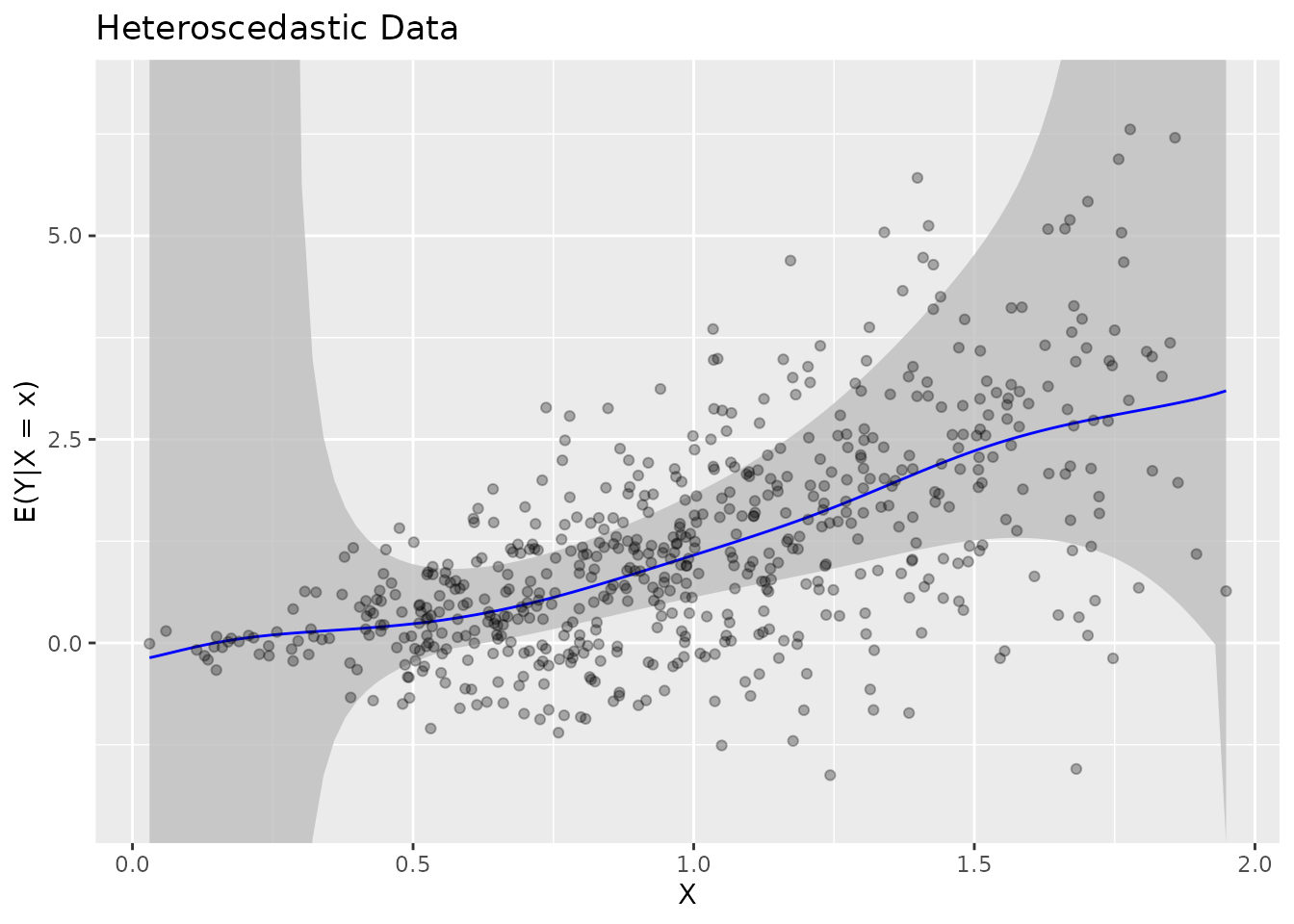
Polynomial Regression Example
# Quadratic relationship
set.seed(456)
X_poly <- gen_sample_data(size = 500, dgp = "2_fold_uniform")
Y_poly <- -X_poly^2 + 3*X_poly + rnorm(500, sd = 0.5)
fit_poly <- biasBound_condExpectation(Y_poly, X_poly, h = 0.1)
# Compare with true function
true_poly <- function(x) -x^2 + 3*x
plot(fit_poly) +
stat_function(fun = true_poly, aes(color = "True: -x^2 + 3x"),
linetype = "dashed", linewidth = 1) +
scale_color_manual(values = c("True: -x^2 + 3x" = "red")) +
labs(color = NULL) +
theme(legend.position = "top")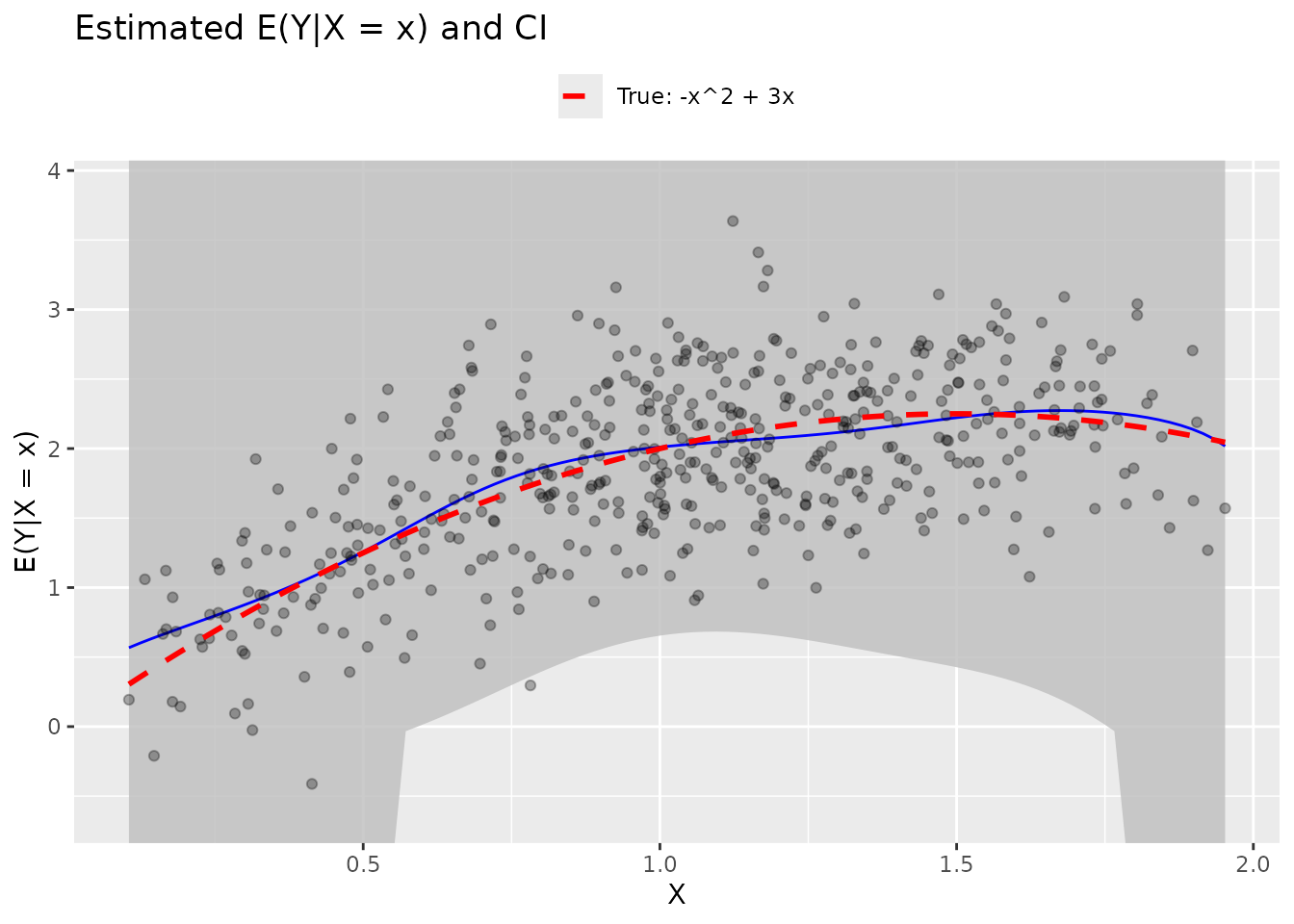
See Also
- Get Started: Quick introduction
- Density Estimation: Density estimation details
- Theory: Mathematical background