Density Estimation with rbbnp
Xinyu Dai and Susanne M. Schennach
2026-02-14
density-estimation.RmdThis article provides a comprehensive guide to density estimation
using the biasBound_density() function.
The biasBound_density() Function
The main function for density estimation takes the following key arguments:
biasBound_density(
X, # Data vector
x = NULL, # Evaluation points (auto if NULL)
h = NULL, # Bandwidth (auto-selected if NULL)
h_method = "cv", # "cv" or "silverman"
alpha = 0.05, # Confidence level
kernel.fun = "Schennach2004",
xi_lb = NULL, # Frequency range lower bound
xi_ub = NULL # Frequency range upper bound
)Basic Example
# Generate sample data from 2-fold uniform distribution
X <- gen_sample_data(size = 1000, dgp = "2_fold_uniform", seed = 42)
# Estimate density
fit <- biasBound_density(X, h = 0.08, kernel.fun = "Schennach2004")
# Display results
fit
#> Bias-Bounded Density Estimation
#>
#> Call:
#> biasBound_density(X = X, h = 0.08, kernel.fun = "Schennach2004")
#>
#> Sample size: n = 1000
#> Bandwidth: h = 0.0800 (user-specified)
#> Kernel: Schennach2004
#>
#> Bias bound parameters:
#> A = 3.4851, r = 1.9202
#> bias bound b1x = 0.1209
#>
#> Evaluation points: 100 (range: [-0.1746, 2.1780])
#> Confidence level: 95%
#>
#> Use summary() for detailed statistics
#> Use plot() to visualize resultsUnderstanding the Output
The result is an S3 object of class bbnp_density
containing:
# Key parameters
coef(fit)
#> A r h
#> 3.485103 1.920169 0.080000
# Detailed summary
summary(fit)
#> Summary: Bias-Bounded Density Estimation
#> ============================================================
#>
#> Call:
#> biasBound_density(X = X, h = 0.08, kernel.fun = "Schennach2004")
#>
#> Sample Information:
#> Sample size (n): 1000
#> Bandwidth (h): 0.0800
#> Kernel function: Schennach2004
#>
#> Bias Bound Parameters:
#> A (amplitude): 3.4851
#> r (decay rate): 1.9202
#> b1x (bias bound): 0.1209
#> Xi interval: [2.4074, 13.5376]
#>
#> Range Estimation:
#> Density estimates:
#> min Q1.25% median mean Q3.75% max
#> 0.0000 0.1111 0.4273 0.4217 0.7125 0.8798
#>
#> Standard errors:
#> min mean max
#> 0.0000 0.0341 0.0563Visualizing Results
Density Plot
plot(fit)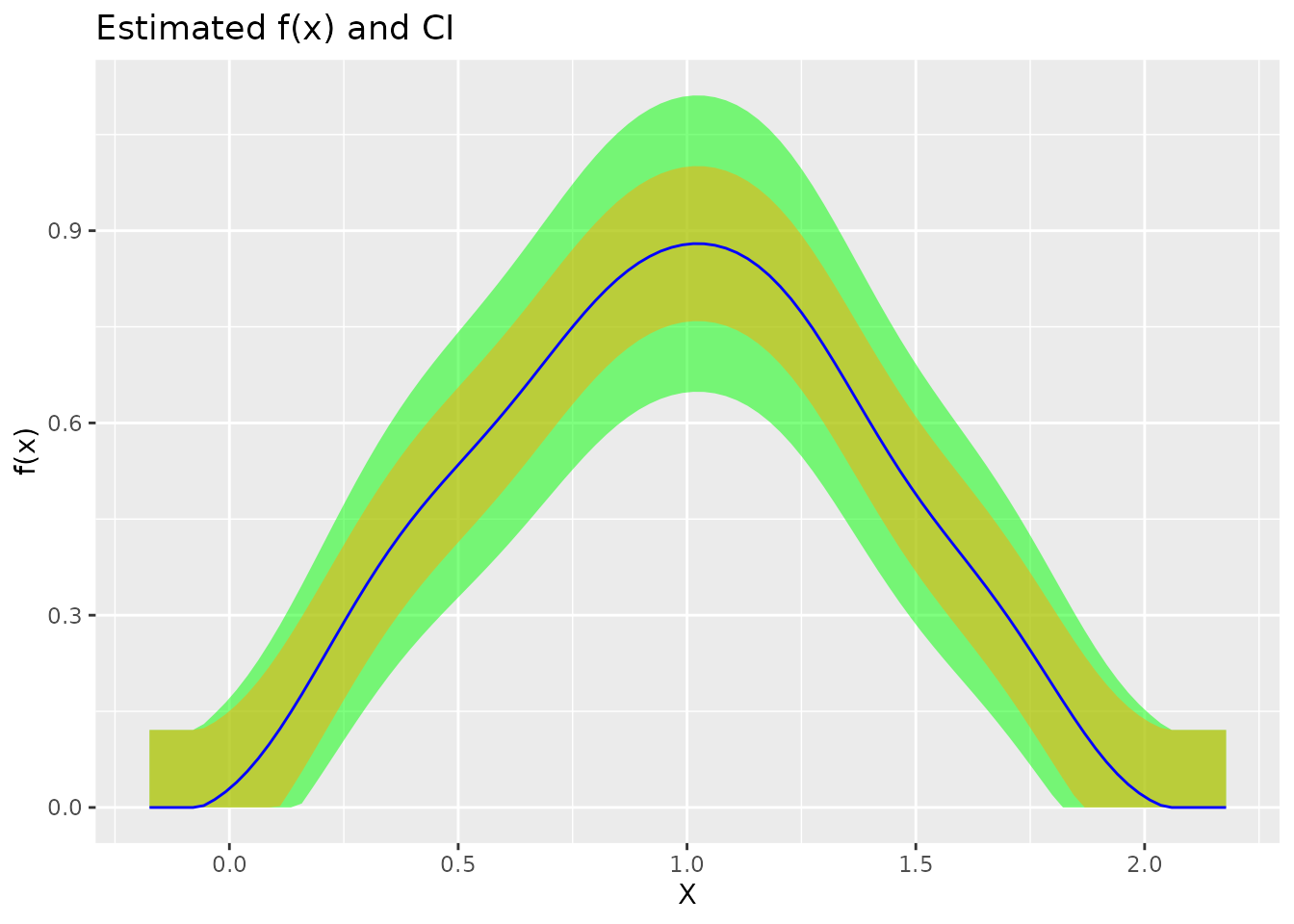
Interpreting the plot:
| Element | Description |
|---|---|
| Blue line | Kernel density estimate |
| Orange band | Bias range: |
| Green band | 95% CI: |
Fourier Transform Analysis
The package automatically detects smoothness via Fourier analysis:
plot(fit, type = "ft")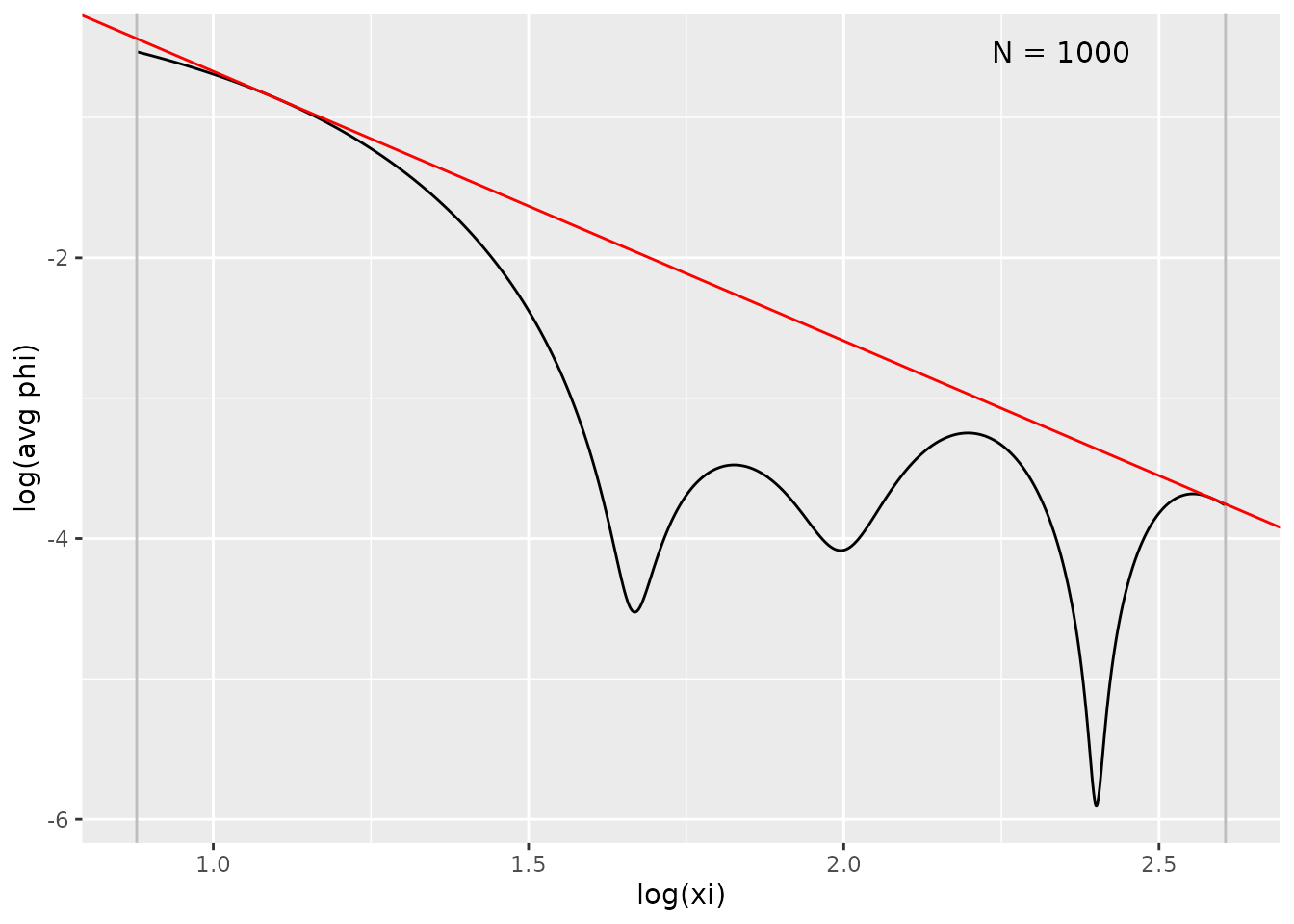
Interpreting the Fourier plot:
- Black curve: Empirical Fourier transform
- Grey lines: Frequency bounds
- Red line: Estimated envelope
The slope indicates smoothness: larger = smoother function.
Bandwidth Selection
Cross-Validation
fit_cv <- biasBound_density(X, h = NULL, h_method = "cv", kernel.fun = "normal")
cat("CV bandwidth:", coef(fit_cv)["h"], "\n")
#> CV bandwidth: 0.1878156
plot(fit_cv) + ggtitle("Cross-Validation Bandwidth")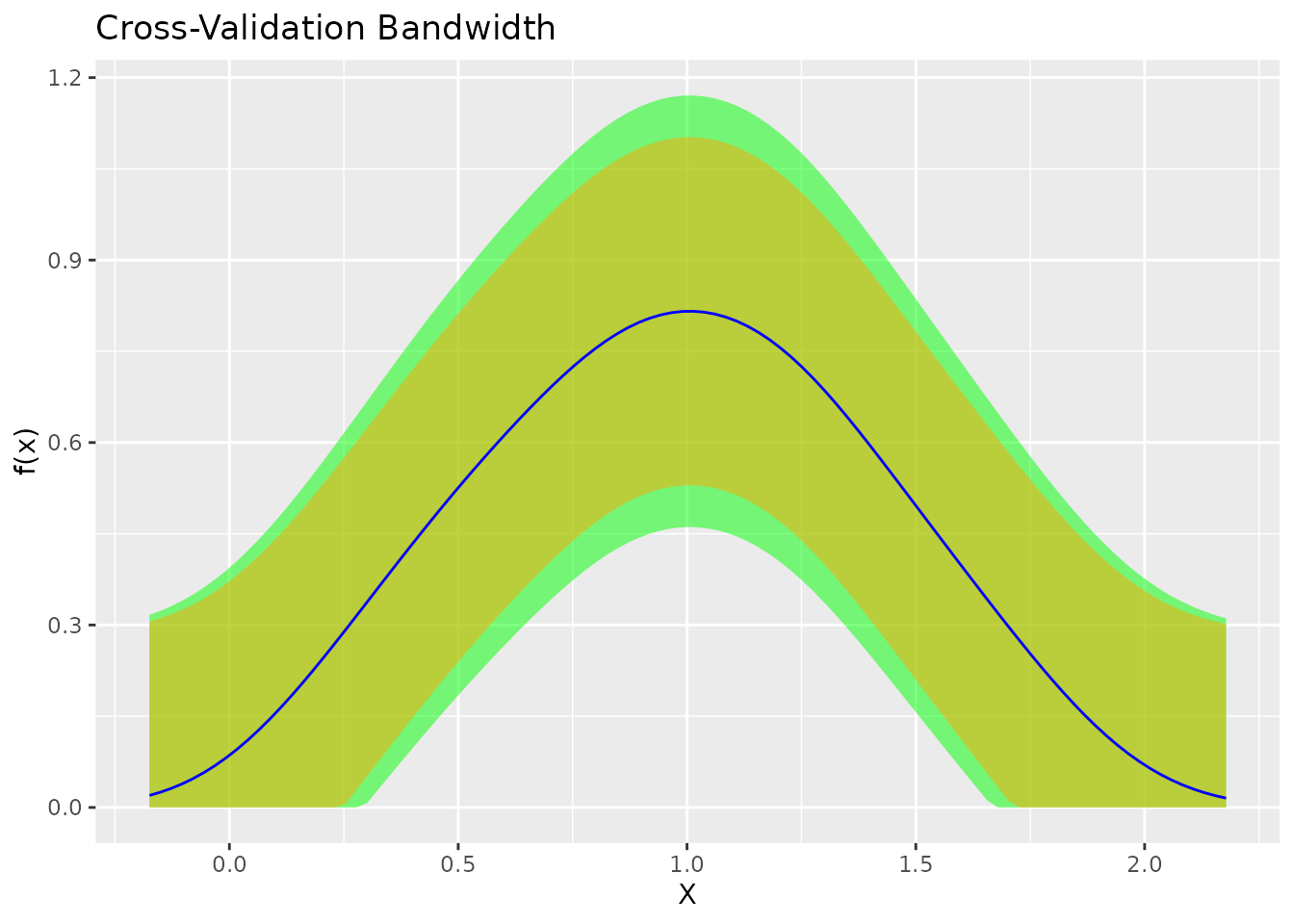
Silverman’s Rule of Thumb
fit_silv <- biasBound_density(X, h = NULL, h_method = "silverman", kernel.fun = "normal")
cat("Silverman bandwidth:", coef(fit_silv)["h"], "\n")
#> Silverman bandwidth: 0.09390781
plot(fit_silv) + ggtitle("Silverman's Rule Bandwidth")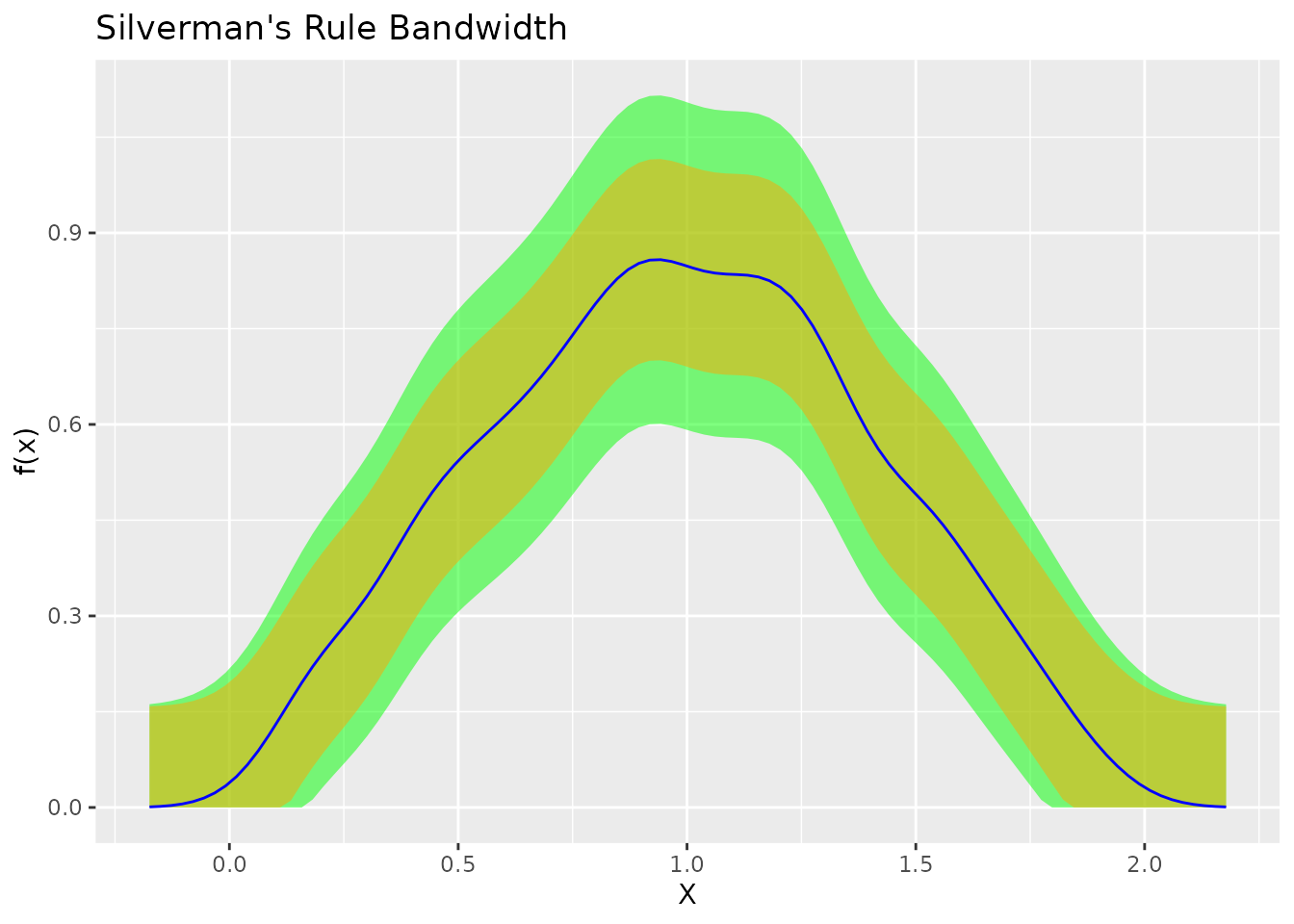
Direct Bandwidth Selection
# Select bandwidth without full estimation
h_cv <- select_bandwidth(X, method = "cv", kernel.fun = "normal")
h_silv <- select_bandwidth(X, method = "silverman", kernel.fun = "normal")
cat("CV:", h_cv, "\nSilverman:", h_silv)
#> CV: 0.1878156
#> Silverman: 0.09390781Kernel Functions
Available Kernels
| Kernel | Order | Best For |
|---|---|---|
| Schennach2004 | Infinite | High smoothness (recommended) |
| sinc | Infinite | Any smoothness level |
| normal | 2 | Moderate smoothness |
| epanechnikov | 2 | Limited smoothness |
Comparing Kernels
fit_sch <- biasBound_density(X, kernel.fun = "Schennach2004")
fit_sinc <- biasBound_density(X, kernel.fun = "sinc")
fit_norm <- biasBound_density(X, kernel.fun = "normal")
fit_epan <- biasBound_density(X, kernel.fun = "epanechnikov")
grid.arrange(
plot(fit_sch) + ggtitle("Schennach2004 (infinite order)"),
plot(fit_sinc) + ggtitle("Sinc (infinite order)"),
plot(fit_norm) + ggtitle("Normal (2nd order)"),
plot(fit_epan) + ggtitle("Epanechnikov (2nd order)"),
ncol = 2
)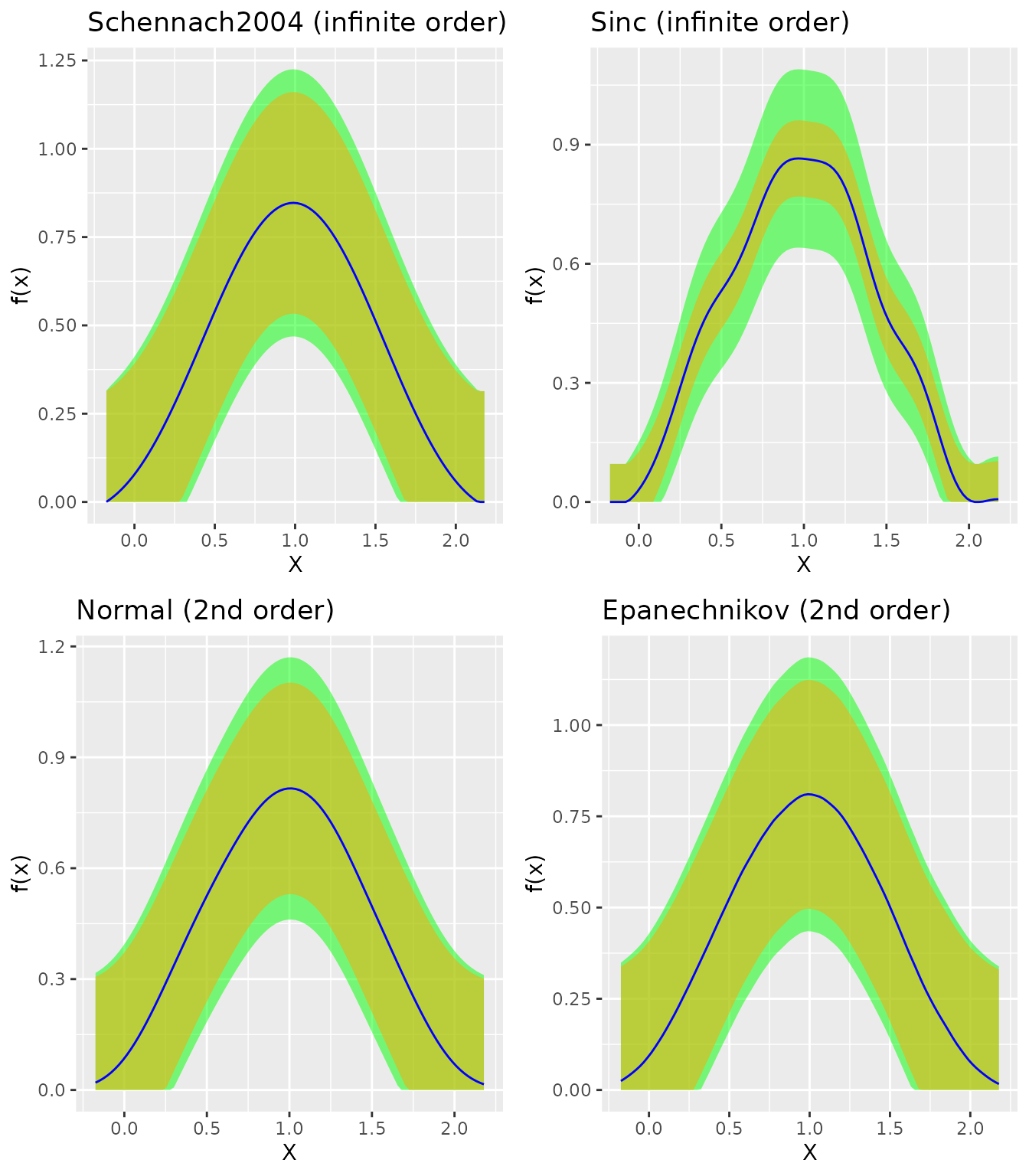
Recommendation: Use "Schennach2004" for
most applications as it adapts to any smoothness level.
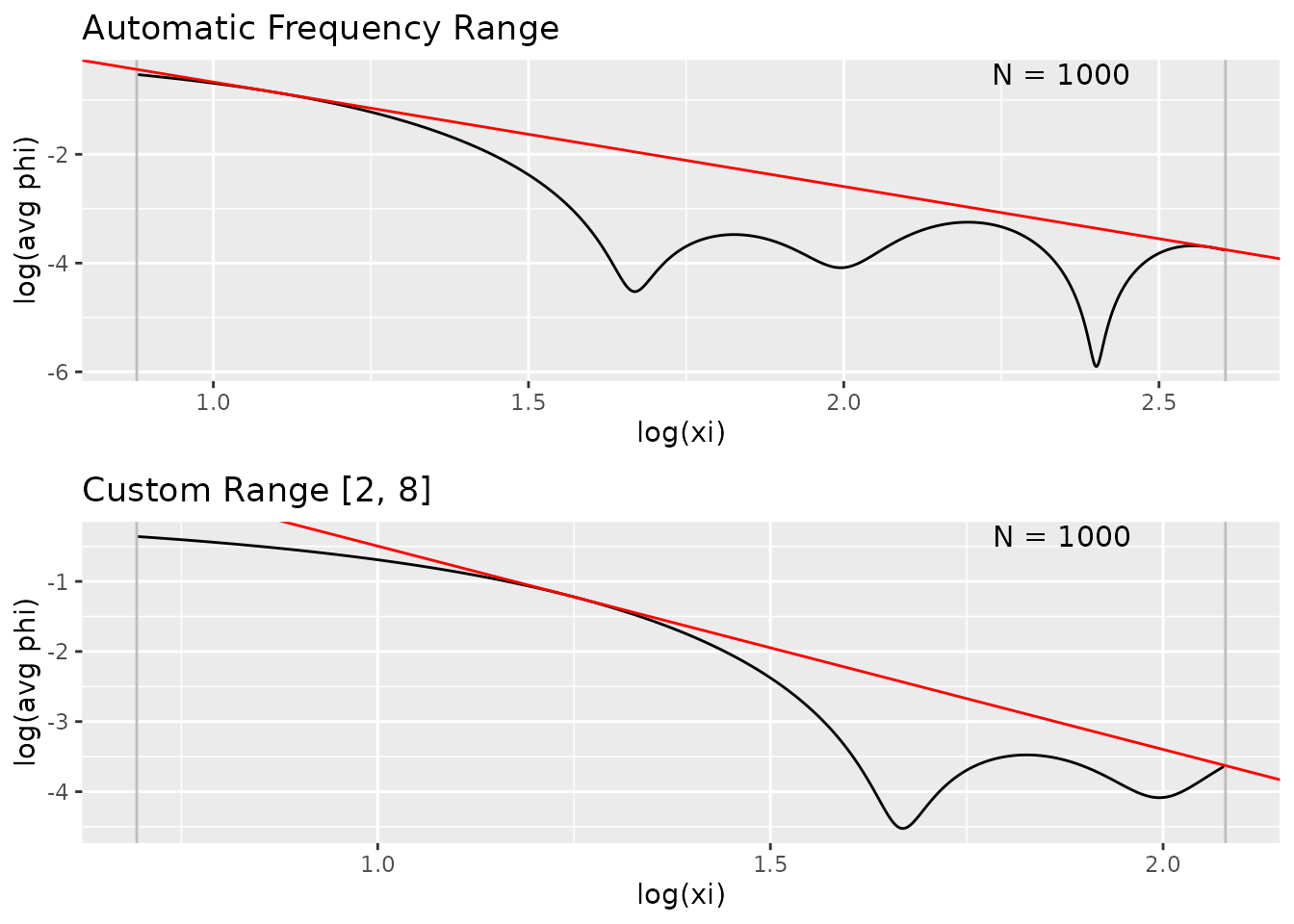
Custom Frequency Range
You can manually specify the frequency range for Fourier analysis:
# Default (automatic)
fit_auto <- biasBound_density(X, h = 0.08)
# Custom range
fit_custom <- biasBound_density(X, h = 0.08, xi_lb = 2, xi_ub = 8)
grid.arrange(
plot(fit_auto, type = "ft") + ggtitle("Automatic Frequency Range"),
plot(fit_custom, type = "ft") + ggtitle("Custom Range [2, 8]"),
ncol = 1
)
Extracting Results
# Confidence intervals as matrix
ci <- confint(fit)
head(ci, 10)
#> lower upper
#> [1,] 0 0.1209374
#> [2,] 0 0.1209374
#> [3,] 0 0.1209374
#> [4,] 0 0.1209374
#> [5,] 0 0.1209374
#> [6,] 0 0.1302624
#> [7,] 0 0.1464837
#> [8,] 0 0.1637745
#> [9,] 0 0.1833616
#> [10,] 0 0.2053695
# Evaluation points
x_points <- fit$x
head(x_points)
#> [1] -0.1746134 -0.1508492 -0.1270850 -0.1033208 -0.0795566 -0.0557924
# Density estimates
f_hat <- fit$f_hat
head(f_hat)
#> NULLComparison with Undersmoothing
The bias-bound approach yields narrower CIs than undersmoothing:
h_opt <- select_bandwidth(X, method = "cv", kernel.fun = "sinc")
# Bias-bound with optimal bandwidth
result_opt <- biasBound_density(X, h = h_opt, kernel.fun = "sinc")
# Undersmoothing (half optimal)
result_under <- biasBound_density(X, h = h_opt * 0.5, kernel.fun = "sinc")
grid.arrange(
plot(result_opt) + ggtitle(paste0("Bias Bound (h = ", round(h_opt, 3), ")")),
plot(result_under) + ggtitle(paste0("Undersmoothing (h = ", round(h_opt/2, 3), ")")),
ncol = 1
)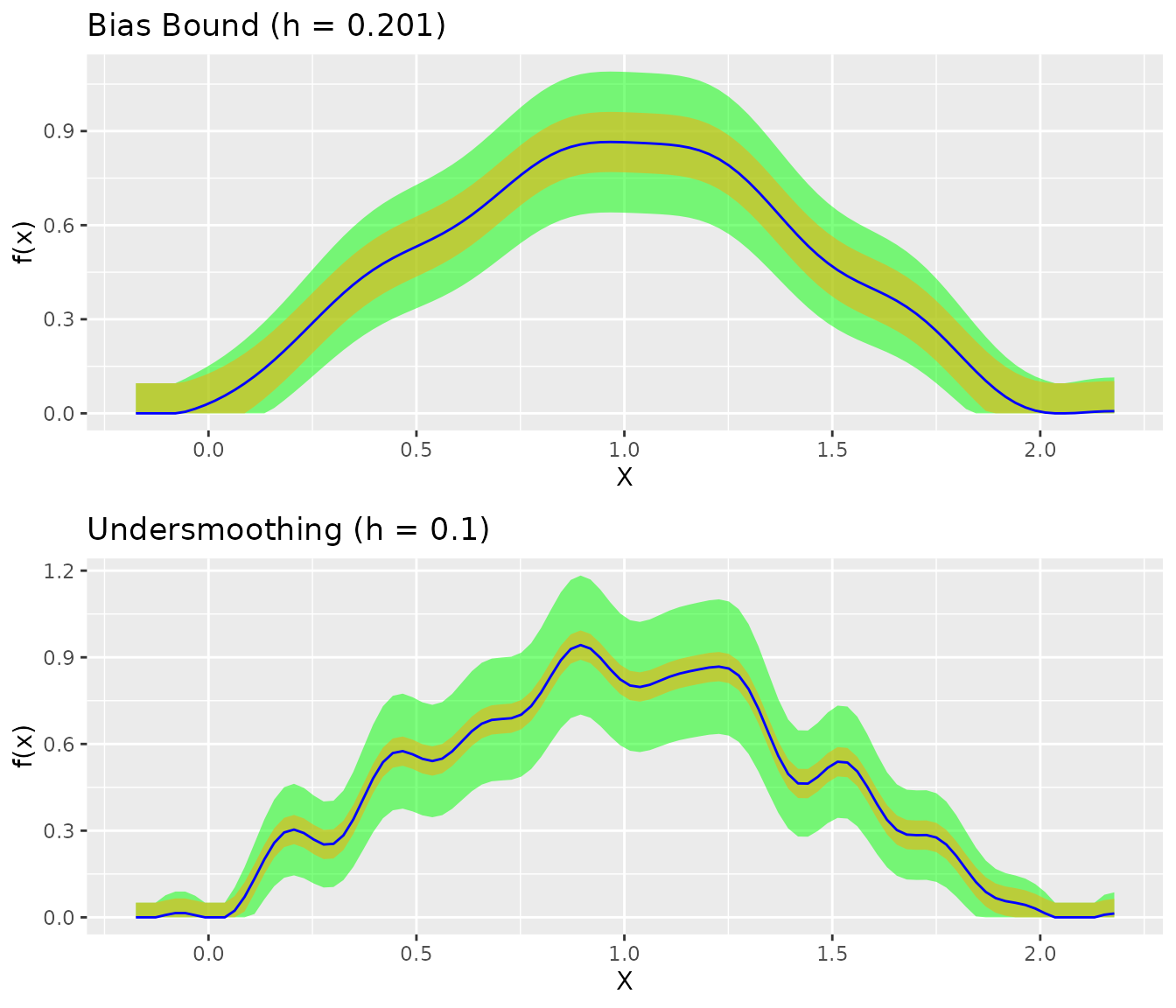
The bias-bound approach provides valid coverage while maintaining efficient estimation.
See Also
- Get Started: Quick introduction
- Regression: Conditional expectation estimation
- Theory: Mathematical background